교재 : 파이썬 머신러닝 완벽 가이드, 위키북스
결정 트리
결정 트리 실습
Human Activity Recognition Using Smartphones Data Set
*https://archive.ics.uci.edu/ml/datasets/human+activity+recognition+using+smartphones
import pandas as pd
import matplotlib.pyplot as pltpd.read_csv('human_activity/features.txt',
sep='\s+',
header=None,
names=['column_index','column_name'])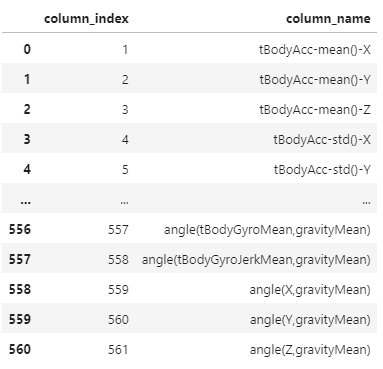
def get_new_df(old_df):
dup_df = pd.DataFrame(data=old_df.groupby('column_name').cumcount(),columns=['dup_cnt'])
new_df = dup_df.reset_index()
return new_dfdf = get_new_df(feature_name_df)
df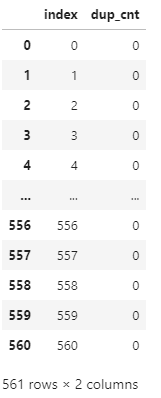
def get_new_df(old_df):
dup_df = pd.DataFrame(data=old_df.groupby('column_name').cumcount(),columns=['dup_cnt'])
dup_df = dup_df.reset_index()
new_df = pd.merge(old_df.reset_index(),dup_df)
return new_dfdf = get_new_df(feature_name_df)
df
def get_new_df(old_df):
dup_df = pd.DataFrame(data=old_df.groupby('column_name').cumcount(),columns=['dup_cnt'])
dup_df = dup_df.reset_index()
new_df = pd.merge(old_df.reset_index(),dup_df,how='outer')
new_df['column_name'] = new_df[['column_name','dup_cnt']].apply(lambda x: x[0]+'_'+str(x[1]) if x[1]>0 else x[0],axis=1)
new_df.drop(columns=['index'],inplace=True)
return new_dfdf = get_new_df(feature_name_df)
df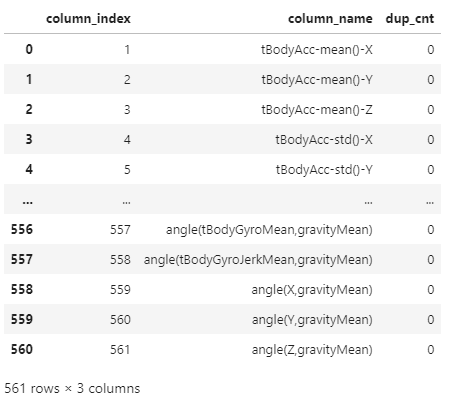
def get_human_dataset():
feature_name_df = pd.read_csv('human_activity/features.txt',
sep='\s+',
header=None,
names=['column_index','column_name'])
name_df = get_new_df(feature_name_df)
feature_name = name_df.iloc[:,1].values.tolist()
X_train = pd.read_csv('human_activity/train/X_train.txt',sep='\s+',names=feature_name)
X_test = pd.read_csv('human_activity/test/X_test.txt',sep='\s+',names=feature_name)
y_train = pd.read_csv('human_activity/train/y_train.txt',sep='\s+',names=['action'])
y_test = pd.read_csv('human_activity/test/y_test.txt',sep='\s+',names=['action'])
return X_train,X_test,y_train,y_testX_train,X_test,y_train,y_test = get_human_dataset()X_train.head(2)
from sklearn.tree import DecisionTreeClassifier
from sklearn.metrics import accuracy_scoredt_clf = DecisionTreeClassifier(random_state=156)
dt_clf.fit(X_train,y_train)
pred = dt_clf.predict(X_test)
accuracy_score(y_test,pred)0.8547675602307431cv_result[['param_max_depth','mean_test_score']]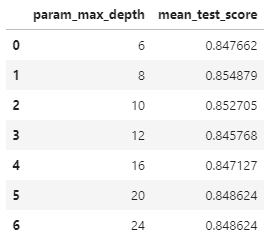
max_depth = [6,8,10,12,16,20,24]
for depth in max_depth:
dt_clf = DecisionTreeClassifier(max_depth=depth,min_samples_split=16,random_state=156)
dt_clf.fit(X_train,y_train)
pred = dt_clf.predict(X_test)
accuracy = accuracy_score(y_test,pred)
print(f'max_depth:{depth} 정확도:{accuracy:.4f}')max_depth:6 정확도:0.8551
max_depth:8 정확도:0.8717
max_depth:10 정확도:0.8599
max_depth:12 정확도:0.8571
max_depth:16 정확도:0.8599
max_depth:20 정확도:0.8565
max_depth:24 정확도:0.8565%%time
params = {
'max_depth': [8,12,16,20],
'min_samples_split': [16,24],
}
grid_cv = GridSearchCV(dt_clf,params,scoring='accuracy',cv=5,verbose=1)
grid_cv.fit(X_train,y_train)Fitting 5 folds for each of 8 candidates, totalling 40 fits
Wall time: 1min 20s
GridSearchCV(cv=5,
estimator=DecisionTreeClassifier(max_depth=24,
min_samples_split=16,
random_state=156),
param_grid={'max_depth': [8, 12, 16, 20],
'min_samples_split': [16, 24]},
scoring='accuracy', verbose=1)grid_cv.best_params_{'max_depth': 8, 'min_samples_split': 16}grid_cv.best_score_0.8548794147162603%%time
params = {
'max_depth': [8],
'min_samples_split': [8,12,16,20,24],
}
grid_cv = GridSearchCV(dt_clf,params,scoring='accuracy',cv=5,verbose=1)
grid_cv.fit(X_train,y_train)Fitting 5 folds for each of 5 candidates, totalling 25 fitsgrid_cv.best_params_grid_cv.best_score_pred = grid_cv.best_estimator_.predict(X_test)
accuracy_score(y_test,pred)grid_cv.best_estimator_.feature_importances_pd.Series(grid_cv.best_estimator_.feature_importances_)data = pd.Series(grid_cv.best_estimator_.feature_importances_,index=X_train.columns)data.sort_values(ascending=False)top10 = data.sort_values(ascending=False)[:10]import seaborn as snssns.barplot(x=top10,y=top10.index)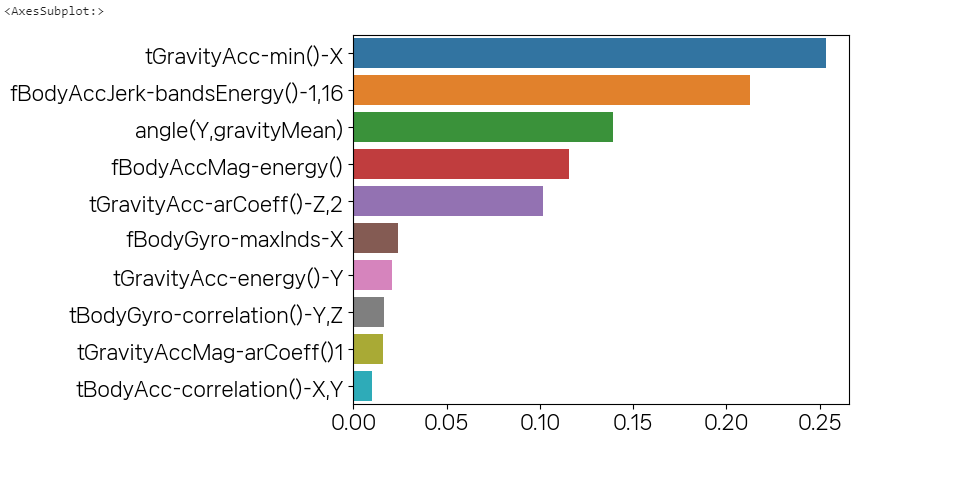
앙상블 학습(Ensemble Learning)
- 비정형은 딥러닝이 뛰어나지만, 정형 데이터 분류와 회귀 분야에서는 앙상블을 사용함
- 보팅, 배깅, 부스팅이 대표적인 앙상블 유형
- 보팅 : 하나의 데이터 셋을 여러 모델로 돌려 투표로 예측함
- 배깅 : 여러 개의 데이터 셋을 하나의 모델로 돌려 투표로 예측함
- 부스팅: 여러 분류기로 순차적으로 학습. 예측을 맞춘 데이터에 가중치를 부여함.
보팅
- 하드 보팅 : 분류기 투표의 다수결로 최종 레이블 값을 결정
- 소프트 보팅 : 분류기 결과물의 평균값으로 결정
- 일반적으로는 소프트 보팅의 예측 성능이 더 좋다.
보팅 분류기(Voting Classifier)
import pandas as pd
from sklearn.ensemble import VotingClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.neighbors import KNeighborsClassifier
from sklearn.datasets import load_breast_cancer
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_scorecancer = load_breast_cancer(as_frame=True)cancer.target.head(2)0 0
1 0
Name: target, dtype: int32cancer.target_namesarray(['malignant', 'benign'], dtype='<U9')lr_clf = LogisticRegression(solver='liblinear')
knn_clf = KNeighborsClassifier(n_neighbors=8)
vo_clf = VotingClassifier([('lr',lr_clf),('knn',knn_clf)],voting='soft')
X_train,X_test,y_train,y_test = train_test_split(cancer.data,
cancer.target,
test_size=0.2,
random_state=156)
vo_clf.fit(X_train,y_train)
pred = vo_clf.predict(X_test)
accuracy_score(y_test,pred)0.956140350877193models = [lr_clf,knn_clf]
for model in models:
model.fit(X_train,y_train)
pred = model.predict(X_test)
model_name = model.__class__.__name__
print(f'{model_name} 정확도 : {accuracy_score(y_test,pred)}')LogisticRegression 정확도 : 0.9473684210526315
KNeighborsClassifier 정확도 : 0.9385964912280702랜덤 포레스트
- 배깅 방식을 이용
- 전체 데이터에서 여러 개의 표본 데이터를 추출 후 같은 모델로 학습
- 단점 : 하이퍼 파라미터가 많고 튜닝이 오래 걸림. 튜닝에 많은 시간을 써도 성능이 잘 향상되지 않음.
from sklearn.ensemble import RandomForestClassifierdef get_new_df(old_df):
dup_df = pd.DataFrame(data=old_df.groupby('column_name').cumcount(),columns=['dup_cnt'])
dup_df = dup_df.reset_index()
new_df = pd.merge(old_df.reset_index(),dup_df,how='outer')
new_df['column_name'] = new_df[['column_name','dup_cnt']].apply(lambda x: x[0]+'_'+str(x[1]) if x[1]>0 else x[0],axis=1)
new_df.drop(columns=['index'],inplace=True)
return new_df
def get_human_dataset():
feature_name_df = pd.read_csv('human_activity/features.txt',
sep='\s+',
header=None,
names=['column_index','column_name'])
name_df = get_new_df(feature_name_df)
feature_name = name_df.iloc[:,1].values.tolist()
X_train = pd.read_csv('human_activity/train/X_train.txt',sep='\s+',names=feature_name)
X_test = pd.read_csv('human_activity/test/X_test.txt',sep='\s+',names=feature_name)
y_train = pd.read_csv('human_activity/train/y_train.txt',sep='\s+',names=['action'])
y_test = pd.read_csv('human_activity/test/y_test.txt',sep='\s+',names=['action'])
return X_train,X_test,y_train,y_testX_train,X_test,y_train,y_test = get_human_dataset()rf_clf = RandomForestClassifier(random_state=0,max_depth=8)
rf_clf.fit(X_train,y_train)
pred = rf_clf.predict(X_test)
accuracy_score(y_test,pred)C:\Users\user\AppData\Local\Temp\ipykernel_18236\990638652.py:2: DataConversionWarning: A column-vector y was passed when a 1d array was expected. Please change the shape of y to (n_samples,), for example using ravel().
rf_clf.fit(X_train,y_train)
0.9195792331184255rf_clf = RandomForestClassifier(random_state=0)
rf_clf.fit(X_train,y_train)
pred = rf_clf.predict(X_test)
accuracy_score(y_test,pred)C:\Users\user\AppData\Local\Temp\ipykernel_18236\1442801055.py:2: DataConversionWarning: A column-vector y was passed when a 1d array was expected. Please change the shape of y to (n_samples,), for example using ravel().
rf_clf.fit(X_train,y_train)
0.9253478113335596from sklearn.model_selection import GridSearchCVparams ={
'max_depth':[8,16,24],
'min_samples_split':[2,8,16],
'min_samples_leaf':[1,6,12]
}%%time
rf_clf = RandomForestClassifier(random_state=0,n_jobs=-1)
grid_cv = GridSearchCV(rf_clf,params,cv=2,n_jobs=-1)
grid_cv.fit(X_train,y_train)C:\Users\user\anaconda3\lib\site-packages\sklearn\model_selection\_search.py:926: DataConversionWarning: A column-vector y was passed when a 1d array was expected. Please change the shape of y to (n_samples,), for example using ravel().
self.best_estimator_.fit(X, y, **fit_params)
Wall time: 36.6 s
GridSearchCV(cv=2, estimator=RandomForestClassifier(n_jobs=-1, random_state=0),
n_jobs=-1,
param_grid={'max_depth': [8, 16, 24],
'min_samples_leaf': [1, 6, 12],
'min_samples_split': [2, 8, 16]})grid_cv.best_params_{'max_depth': 16, 'min_samples_leaf': 6, 'min_samples_split': 2}grid_cv.best_score_0.9164853101196953rf_clf = RandomForestClassifier(random_state=0,max_depth=16,min_samples_leaf=6,min_samples_split=2)
rf_clf.fit(X_train,y_train)
pred = rf_clf.predict(X_test)
accuracy_score(y_test,pred)C:\Users\user\AppData\Local\Temp\ipykernel_18236\795056095.py:2: DataConversionWarning: A column-vector y was passed when a 1d array was expected. Please change the shape of y to (n_samples,), for example using ravel().
rf_clf.fit(X_train,y_train)
0.9260264675941635부스팅(Boosting)
GBM(Gradient Boosting Machine)
- 에이다부스트와 그래디언트 부스트가 있음
- 경사하강법을 이용하여 오류 최소화로 가중치를 업데이트함
import pandas as pd
from sklearn.ensemble import GradientBoostingClassifier
import warnings
warnings.filterwarnings('ignore')
from sklearn.metrics import accuracy_scoredef get_new_df(old_df):
dup_df = pd.DataFrame(data=old_df.groupby('column_name').cumcount(),columns=['dup_cnt'])
dup_df = dup_df.reset_index()
new_df = pd.merge(old_df.reset_index(),dup_df,how='outer')
new_df['column_name'] = new_df[['column_name','dup_cnt']].apply(lambda x: x[0]+'_'+str(x[1]) if x[1]>0 else x[0],axis=1)
new_df.drop(columns=['index'],inplace=True)
return new_df
def get_human_dataset():
import pandas as pd
feature_name_df = pd.read_csv('human_activity/features.txt',
sep='\s+',
header=None,
names=['column_index','column_name'])
name_df = get_new_df(feature_name_df)
feature_name = name_df.iloc[:,1].values.tolist()
X_train = pd.read_csv('human_activity/train/X_train.txt',sep='\s+',names=feature_name)
X_test = pd.read_csv('human_activity/test/X_test.txt',sep='\s+',names=feature_name)
y_train = pd.read_csv('human_activity/train/y_train.txt',sep='\s+',names=['action'])
y_test = pd.read_csv('human_activity/test/y_test.txt',sep='\s+',names=['action'])
return X_train,X_test,y_train,y_testX_train,X_test,y_train,y_test = get_human_dataset()%%time
gb_clf = GradientBoostingClassifier(random_state=0)
gb_clf.fit(X_train,y_train)
pred = gb_clf.predict(X_test)
accuracy_score(y_test,pred)Wall time: 8min 26s
0.9389209365456397XGBoost(eXtra Gradient Boost)
- 트리 기반 앙상블 학습에서 가장 인기있는 알고리즘
- GBM과 달리 병렬 학습이 가능해 속도가 빠름
XGBoost 설치
import xgboost as xgb
from xgboost import XGBClassifier
from xgboost import plot_importance
import pandas as pd
import numpy as np
from sklearn.datasets import load_breast_cancer
from sklearn.model_selection import train_test_splitxgb.__version__'1.5.0'dataset = load_breast_cancer(as_frame=True)dataset.data
dataset.target0 0
1 0
2 0
3 0
4 0
..
564 0
565 0
566 0
567 0
568 1
Name: target, Length: 569, dtype: int32dataset.target_namesarray(['malignant', 'benign'], dtype='<U9')dataset.target.value_counts()1 357
0 212
Name: target, dtype: int64X_train,X_test,y_train,y_test = train_test_split(dataset.data,dataset.target,test_size=0.2,random_state=156)
X_tr,X_val,y_tr,y_val = train_test_split(X_train,y_train,test_size=0.1,random_state=156)X_train.shape,X_test.shape((455, 30), (114, 30))X_tr.shape,X_val.shape((409, 30), (46, 30))y_train.value_counts()1 280
0 175
Name: target, dtype: int64dtr = xgb.DMatrix(data=X_tr,label=y_tr)
dval = xgb.DMatrix(data=X_val,label=y_val)
dtest = xgb.DMatrix(data=X_test,label=y_test)params = {
'max_depth' : 3,
'eta' : 0.05,
'objective' : 'binary:logistic',
'eval_metric' : 'logloss'
}
num_rounds = 500
eval_list=[(dtr,'train'),(dval,'eval')]xgb.train(params,dtr,num_rounds,evals=eval_list,early_stopping_rounds=50)dataset.target0 0
1 0
2 0
3 0
4 0
..
564 0
565 0
566 0
567 0
568 1
Name: target, Length: 569, dtype: int32dataset.target_namesarray(['malignant', 'benign'], dtype='<U9')dataset.target.value_counts()1 357
0 212
Name: target, dtype: int64X_train,X_test,y_train,y_test = train_test_split(dataset.data,dataset.target,test_size=0.2,random_state=156)
X_tr,X_val,y_tr,y_val = train_test_split(X_train,y_train,test_size=0.1,random_state=156)X_train.shape,X_test.shape((455, 30), (114, 30))X_tr.shape,X_val.shape((409, 30), (46, 30))y_train.value_counts()1 280
0 175
Name: target, dtype: int64dtr = xgb.DMatrix(data=X_tr,label=y_tr)
dval = xgb.DMatrix(data=X_val,label=y_val)
dtest = xgb.DMatrix(data=X_test,label=y_test)params = {
'max_depth' : 3,
'eta' : 0.05,
'objective' : 'binary:logistic',
'eval_metric' : 'logloss'
}
num_rounds = 500
eval_list=[(dtr,'train'),(dval,'eval')]xgb.train(params,dtr,num_rounds,evals=eval_list,early_stopping_rounds=50)np.round(pred_probs[:10],3)array([0.845, 0.008, 0.68 , 0.081, 0.975, 0.999, 0.998, 0.998, 0.996,
0.001], dtype=float32)pred = [1 if x>0.5 else 0 for x in pred_probs]def get_clf_eval(y_test, pred, pred_proba_1):
from sklearn.metrics import accuracy_score, precision_score, recall_score, confusion_matrix, f1_score, roc_auc_score
confusion = confusion_matrix(y_test,pred)
accuracy = accuracy_score(y_test,pred)
precision = precision_score(y_test,pred)
recall = recall_score(y_test,pred)
f1 = f1_score(y_test,pred)
auc = roc_auc_score(y_test,pred_proba_1)
print('오차행렬')
print(confusion)
print(f'정확도:{accuracy:.4f}, 정밀도:{precision:.4f}, 재현율{recall:.4f}, F1:{f1:.4f}, AUC:{auc:.4f}')get_clf_eval(y_test,pred,pred_probs)오차행렬
[[34 3]
[ 2 75]]
정확도:0.9561, 정밀도:0.9615, 재현율0.9740, F1:0.9677, AUC:0.9937plot_importance(model)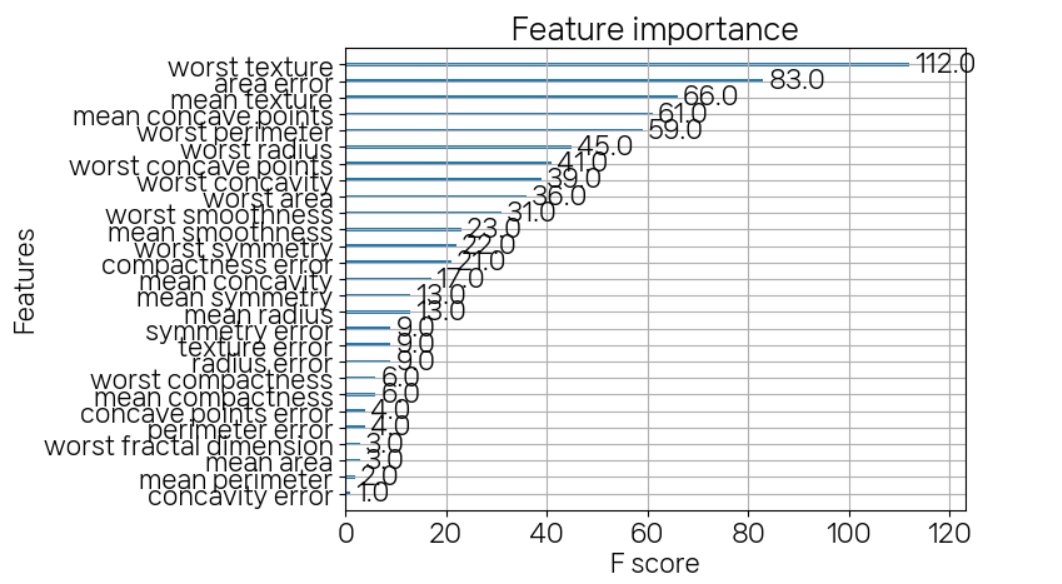
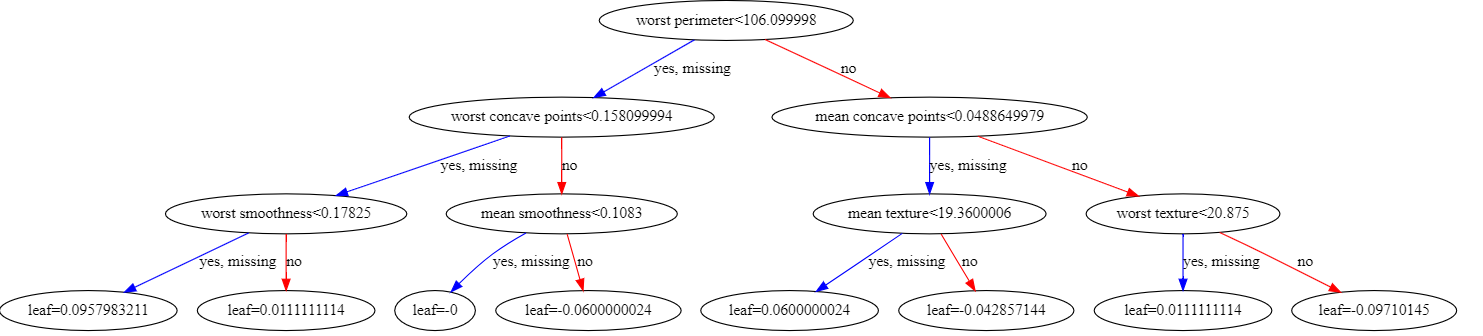
from xgboost import to_graphviz
to_graphviz(model)